SpaRC: Scalable Sequence Clustering using Apache Spark
Автор: insideHPC Report
Загружено: 2018-02-26
Просмотров: 312
In this video from the Stanford HPC Conference, Zhong Wang from the Genome Institute, Lawrence Berkeley National Laboratory presents: SpaRC: Scalable Sequence Clustering using Apache Spark.
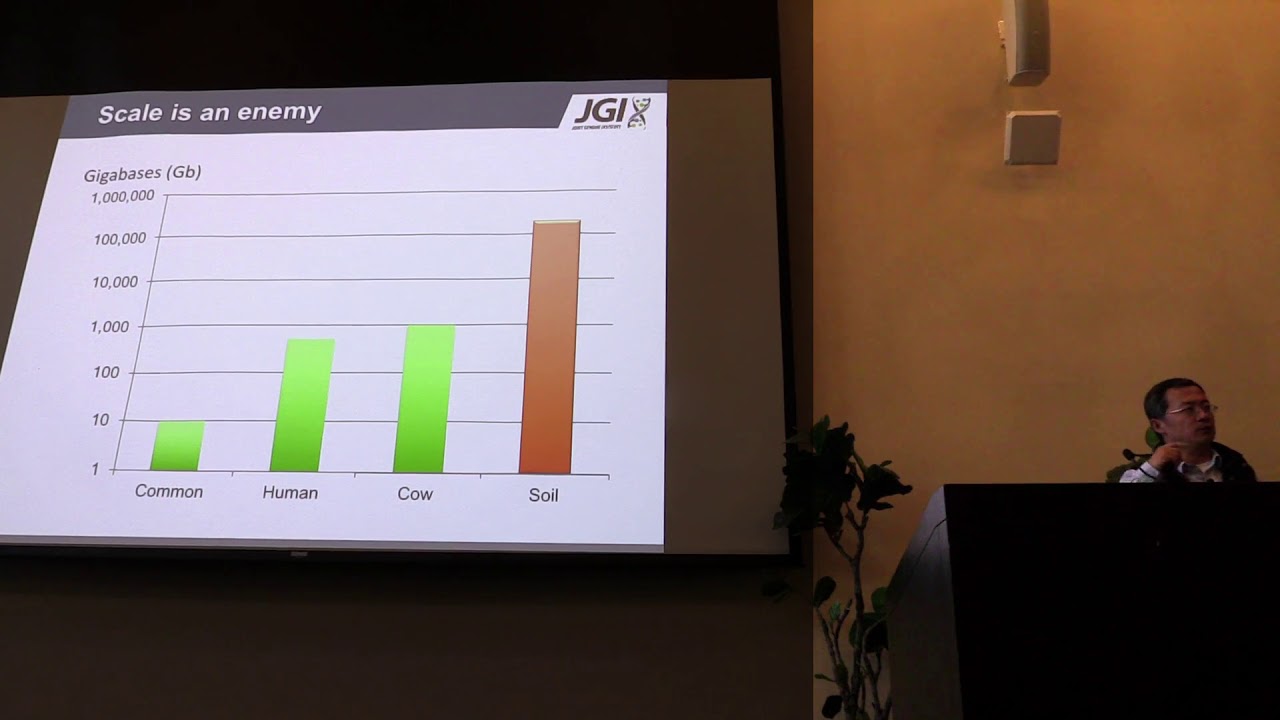
"Whole genome shotgun based next generation transcriptomics and metagenomics studies often generate 100 to 1000 gigabytes (GB) sequence data derived from tens of thousands of different genes or microbial species. Assembly of these data sets requires tradeoffs between scalability and accuracy. Current assembly methods optimized for scalability often sacrifice accuracy and vice versa. An ideal solution would both scale and produce optimal accuracy for individual genes or genomes. Here we describe an Apache Spark-based scalable sequence clustering application, SparkReadClust (SpaRC) that partitions reads based on their molecule of origin to enable downstream assembly optimization. SpaRC produces high clustering performance on transcriptomes and metagenomes from both short and long read sequencing technologies. It achieves near linear scalability with input data size and number of compute nodes. SpaRC can run on different cloud computing environments without modification while delivering similar performance. Our results demonstrate that SpaRC provides a scalable solution for clustering billions of reads from next-generation sequencing experiments, and Apache Spark represents a cost-effective solution with rapid development/deployment cycles for similar large scale sequence data analysis problems. The software is available under the BSD license at https://bitbucket.org/LizhenShi/sparc."
Learn more: https://bitbucket.org/LizhenShi/sparc
and
http://hpcadvisorycouncil.com
Sign up for our insideHPC Newsletter: http://insidehpc.com/newsletter
Доступные форматы для скачивания:
Скачать видео mp4
-
Информация по загрузке: